
GenerateReports
GenerateReports: sequencing data becomes clear and actionable
While new DNA sequencing technologies make it possible to better characterize tumors, they generate a large volume of data that must be interpreted.
GenerateReports is a bioinformatics platform for analyzing raw next-generation sequencing data into a clinical report that clearly summarizes key sequencing results.
For each sample sequenced, there is a report showing :
– sample identification data
– Run-wide quality control
– a specific quality control of the sample
– the list of genetic variations detected and annotated
– variations in the number of gene copies
– experimental traceability and bioinformatics traceability information.


Data aggregation
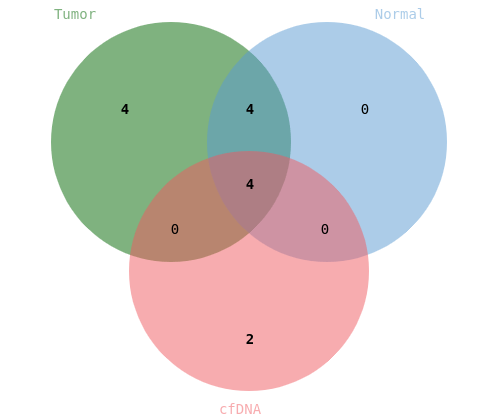
All the data is also aggregated in a database allowing the retrospective consultation of the results at the scale:
– of a complete project
– from a cohort of patients
– an experiment
– a sample or a variant.
Variation lists can be created by biologists, making it possible to create documents, custom filters and certain automated analyses.
The database also allows the storage of external experimental validations to confirm the results obtained by high-throughput sequencing.
The software allows the insertion of data from various sequencing platforms.
Simplicity and efficiency
Its design allows:
- interpretation of samples on a daily basis as part of a diagnostic activity
- a retrospective analysis of biological samples as part of research projects
GenerateReports is above all a modular tool allowing the creation of bioinformatics processing chains according to the nature of the constructions of the sequencing libraries (targeted panels, exomes, capture, amplicon, etc.) and the biological questions asked (diagnosis/research, tumor/ctDNA sequencing…).
It is a help to accreditation for the diagnosis of high-throughput sequencing technology (NGS) by allowing fine control of the versions of bioinformatics pipelines and their dependencies.
GenerateReports integrates the concept of list of variants in order to allow personalized bioinformatics analyses.
